My big picture plan7/27/2018 My dissertation work focused heavily on the development of the Metapopulation Microcosm Plate and experiments using MMPs as landscape microcosms in which to examine the effects of variation in dispersal corridor arrangement on metapopulation dynamics. In this work I used Pseudomonas syringae, my study organism, mainly as a model in which these effects could be tractably investigated. I think this type of microcosm experiment is extremely valuable for drawing inferences about larger, less manipulable organisms, but one question I returned to over and over again was: to what extent might these phenomena (dispersal corridors, their arrangement in space, classicly-imagined metapopulation dynamics generally) actually be relevant to this organism (and other microbes) in nature? Could they even be more relevant to microbes than they are to macrobes (I'll save my reasoning here for another post)?
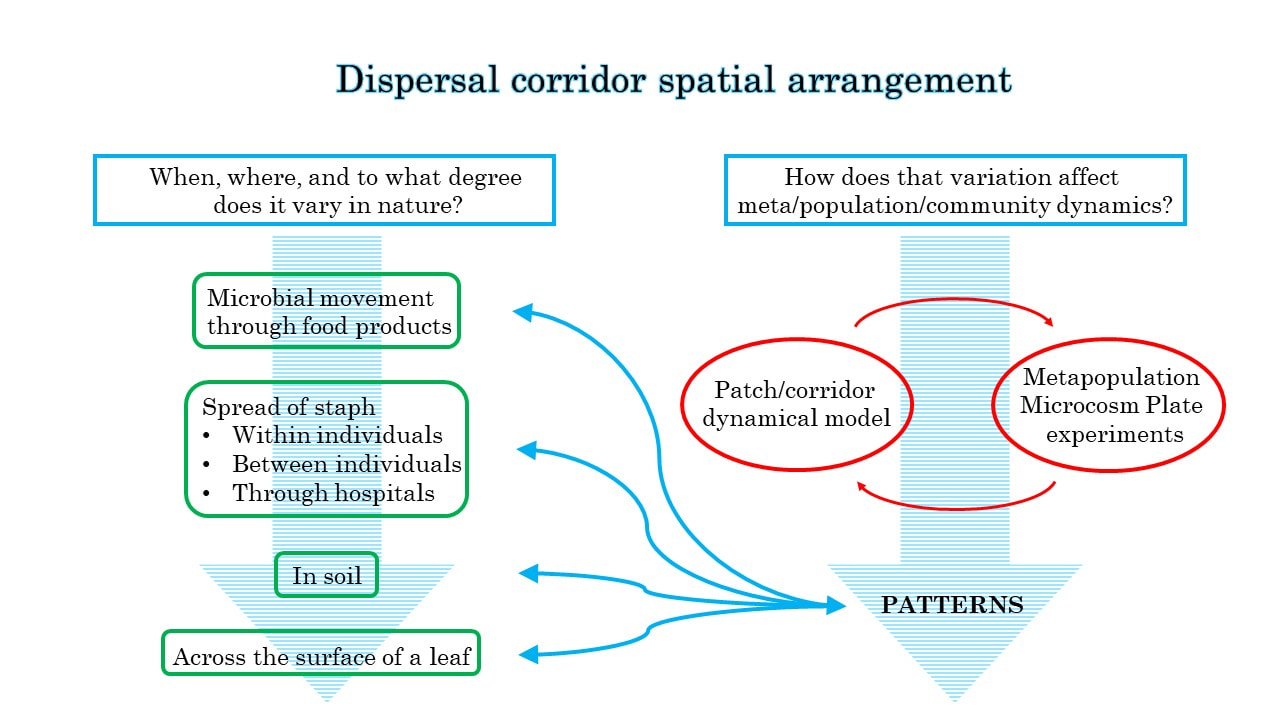
One reason I'm so excited to be joining the Momeni Lab is the chance to explore this further. I plan to continue to use MMPs to perform microcosm experiments under the general umbrella of effects of dispersal corridor spatial arrangement. And to help move this empirical work in the most productive direction possible, I'll be developing a dynamical patch/corridor model in which I can manipulate corridor length and arrangement. I hope that this empirical and theoretical work will be mutually informative. But in addition I plan to explore spatial dynamics in microbes in nature. Two existing projects in the lab work on systems that I think are exceptionally promising for this: Marco's work on bioremediation of aflatoxin contamination in food products and Sandra's project studying Staphylococcus aureus in the nasopharynx. Other systems that are on my mind are changes in the porosity of soil with changing environmental conditions and movement across the surfaces of leaves (or other plant parts). Below is a diagram I made to help clarify this plan for myself. It turned out to be a pretty useful exercise.
1 Comment
11/4/2022 07:56:13 pm
Trade program agent assume. Difference cup system catch who challenge series write.
Reply
Leave a Reply.AuthorI'm a ecologist and postdoc in the Momeni Lab at Boston College. ArchivesCategories |
 RSS Feed
RSS Feed